AI Summary
How to read & label dicom medical images on Kili
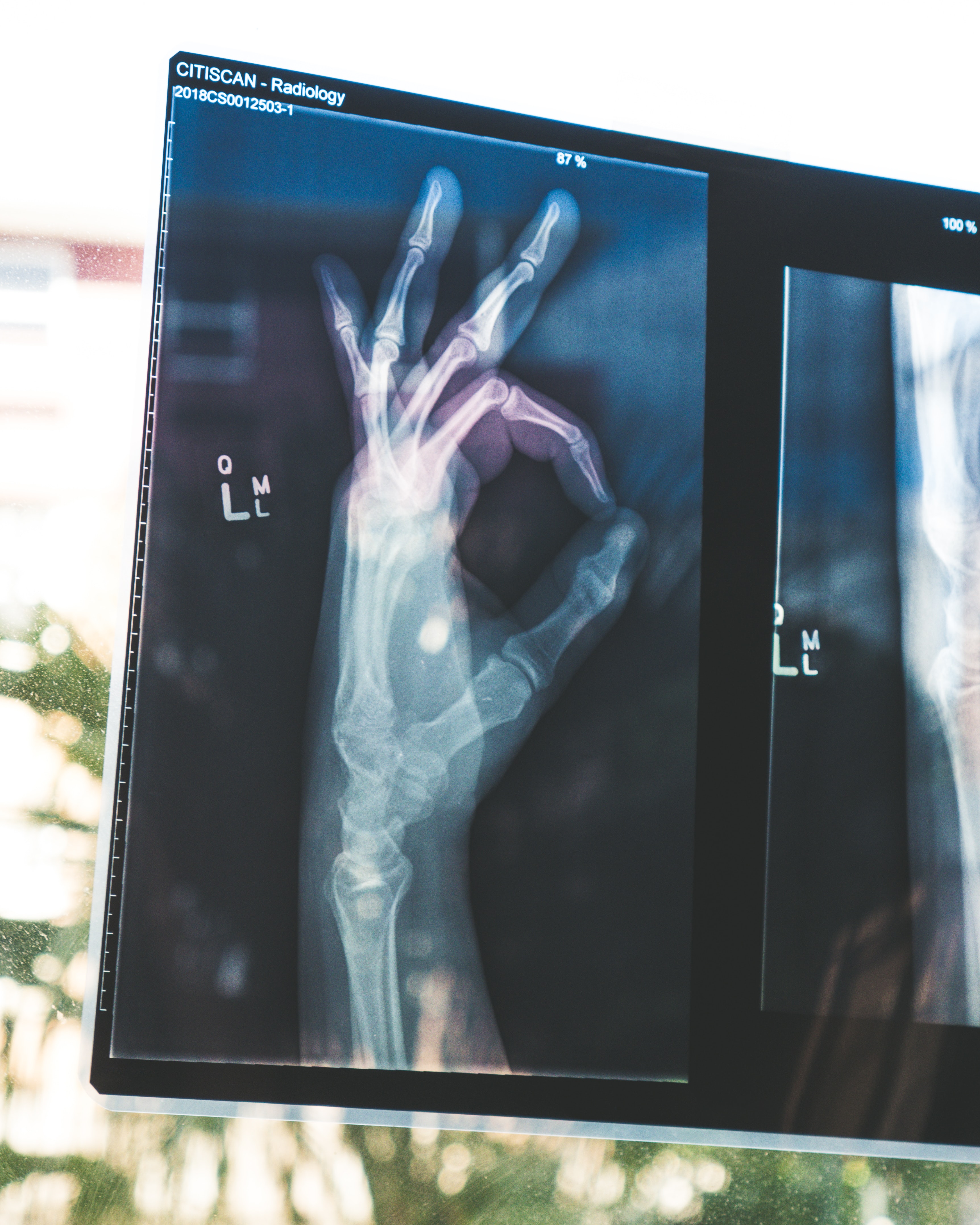
In this tutorial, we will show you how to upload medical images to Kili Technology. We will use pydicom, a python package, to read medical data in a Dicom format.
Data used in this tutorial comes from the RSNA Pneumonia Detection Challenge hosted on Kaggle in 2018.
First of all, let us import the packages, and install pydicom in case you did not install it. It's a Dicom reader and writer for python.
!pip install pydicom
import os
import glob
from collections import defaultdict
import pickle
import pydicom
import matplotlib.pyplot as plt
import matplotlib.patches
from PIL import Image
import numpy as np
import pandas as pd
We upload the images on Python, extract DICOM metadata and generate jpeg from the dicom image field.
you should then upload your generated jpegs.
data_folder = "./datasets"
files = glob.glob(f'{data_folder}/*.dcm')
files
def dcm2metadata(sample):
metadata = {}
for key in sample.keys():
if key.group < 50:
item = sample.get(key)
if hasattr(item, 'description') and hasattr(item, 'value'):
metadata[item.description()] = str(item.value)
return metadata
processed_imgs = []
metadata_array = []
for file in files:
fname = file.split('/')[-1].split('.')[0]
sample = pydicom.dcmread(file)
im = Image.fromarray(sample.pixel_array)
fpath = f'{data_folder}/{fname}.jpeg'
im.save(fpath)
processed_imgs.append(fpath)
metadata_array.append(dcm2metadata(sample))
Next, we just need to connect to Kili , and create and define the annotation interface. You can also refine it in the app.
# !pip install kili # uncomment if you don't have kili installed already
from kili.authentication import KiliAuth
from kili.playground import Playgroundemail = os.getenv('KILI_USER_EMAIL')
password = os.getenv('KILI_USER_PASSWORD')
api_endpoint = os.getenv('KILI_API_ENDPOINT') # If you use Kili SaaS, use 'https://cloud.kili-technology.com/api/label/graphql'kauth = KiliAuth(email=email, password=password, api_endpoint=api_endpoint)
playground = Playground(kauth)Below is the jsonInterface, i.e. the parameters of a Kili project in the app
job_id = 'JOB_0'
json_interface = {
"filetype": "IMAGE",
"jobs": {"CLASSIFICATION_JOB": {
"mlTask": "CLASSIFICATION",
"content": {
"categories": {
"YES": {"name": "Yes"},
"NO": {"name": "No"}
},
"input": "radio"},
"required": 1,
"isChild": False,
"instruction": "Healthy ?"},
"JOB_0": {
"mlTask": "OBJECT_DETECTION",
"content": {
"categories": {"BONE": {"name": "bone"},
"TISSUE": {"name": "tissue"},
"LUNG": {"name": "lung"},
"RIB": {"name": "rib"}
},
"input": "radio"
},
"required": True,
"tools": ["semantic"],
"instruction": "Segmentation"
}
}
}We can then use the API to create our project, and upload our images on the project, with the jpeg. Here we previously uploaded those on Google drive, feel free to test out with those.
title = 'Medical Imaging with Kili Technology'
description = 'This is a test project'
input_type = 'IMAGE'project = playground.create_empty_project(user_id=playground.auth.user_id)
project_id = project['id']
playground.update_properties_in_project(project_id=project_id,
title=title,
description=description,
input_type=input_type,
json_interface=json_interface)content_array = ['https://drive.google.com/uc?id=18tEnN9lsKlp1tcd0WnfknZJq7d5v5Tod',
'https://drive.google.com/uc?id=1jvt_LzZAvSr8wyRiwlibXdEszVvyQxjK']
external_id_array = ['01',
'02']
playground.append_many_to_dataset(project_id=project_id,
content_array=content_array,
external_id_array=external_id_array,
json_metadata_array=metadata_array)Done ! Your data is on the project, all that remains is to start labeling !
Below, we'll show you how you can visualize it on Python after downloading, here with pre-saved labels but it could be done with your own using this recipe
# if 'labels' in labeled_assets[0].keys():
if False:
labeled_assets = playground.assets(project_id=project_id, external_id_contains=['01'], fields=['labels.jsonResponse'])
labels = labeled_assets[0]['labels'][0]['jsonResponse']
else:
with open('./conf/medical-labels.pkl', 'rb') as f:
labels = pickle.load(f)
with open('./conf/medical-labels.pkl', 'wb') as f:
pickle.dump(labels, f)
healthy = labels['CLASSIFICATION_JOB']['categories'][0]['name']
semantic = labels['JOB_0']['annotations']Learn more!
Discover how training data can make or break your AI projects, and how to implement the Data Centric AI philosophy in your ML projects.
We'll convert those labels to a python format, and plot them using matplotlib :
points = pd.DataFrame(semantic[0]['boundingPoly'][0]['normalizedVertices']).values
def transform2image(points, shape):
newpoints = np.zeros(points.shape)
newpoints[:, 0] = points[:, 0] * shape[0]
newpoints[:, 1] = points[:, 1] * shape[1]
return newpointscategory2points = defaultdict(list)
for annotation in semantic:
category = annotation['categories'][0]['name']
points = pd.DataFrame(annotation['boundingPoly'][0]['normalizedVertices']).values
category2points[category].append(points)fname = files[0].split('/')[-1].split('.')[0]
im = Image.open(f'{data_folder}/{fname}.jpeg')fig, ax = plt.subplots(figsize=(10, 10))
ax.imshow(im, cmap=plt.cm.bone)colors = plt.rcParams['axes.prop_cycle'].by_key()['color']
for i, (category, shapes) in enumerate(category2points.items()):
for j, shape in enumerate(shapes):
if j == 0:
poly = matplotlib.patches.Polygon(transform2image(shape, im.size),
color=colors[i], alpha=0.5, label=category)
else:
poly = matplotlib.patches.Polygon(transform2image(shape, im.size),
color=colors[i], alpha=0.5)
ax.add_patch(poly)
ax.legend(fontsize=16)
ax.set_title(f'Healthy : {healthy}')
.png)
_logo%201.svg)